Outbreak Analytics: Data Science Strategies for a Novel Problem
You walk down one aisle of the grocery store to get your favorite cereal. On the dairy aisle, someone sick from COVID-19 coughs. Did your decision to grab your cereal before your milk possibly keep you healthy? How can these unpredictable, near-random choices be included in complex models?
By Susan Sivek, Alteryx.
You walk down one aisle of the grocery store to get your favorite cereal. On the dairy aisle, someone sick from COVID-19 coughs.
Did your decision to grab your cereal before your milk possibly keep you healthy?
It's the kind of question we're all asking as we make simple daily decisions during this global pandemic. But imagine now that it's your job to create a model for the spread of COVID-19 that accounts for humans' unpredictable, near-random choices. Your model also could include government mandates for social distancing, hospital care availability, pre-existing conditions among a population, and more.
Sound complicated? It sure is, but researchers in a variety of fields are working to create the most accurate, useful models possible to predict and explain the spread of COVID-19. I'm not an epidemiologist, but if you too have been impressed and intrigued by the data visualizations and modeling publicized so far, this post is for you.
I explored some recent research to learn a little bit about how these researchers go about what some call outbreak analytics. Here's an introductory look at just a few components of this unique analytics process. We'll find some lessons for other analytics applications, too.
Estimating Virus Transmission
"Once we know the R0, we'll be able to get a handle on the scale of the epidemic."
- the movie Contagion
The reproduction number, or R0 (pronounced "R naught"), refers to the number of people someone who is ill is likely to infect. The R0 changes in every time and place that a disease occurs due to factors like demographics, climate, social structures, and social distancing measures. Researchers calculate the R0 by determining the time between a first infection and a second infection (the generation time, which can then be charted as a generation time distribution when this time is known for many pairs of sick people). Estimating the R0 is critical to predicting a disease's spread. For SARS-CoV-2, which causes COVID-19, the R0 is currently thought to vary between 1.5 to 3.5 in different places.
An R package called R0 (available here) can calculate the current R0 using outbreak data. Offering five different methods for calculation, it also has an option for sensitivity analysis that shows which selection of time window or generation time provided the best fit to the data.
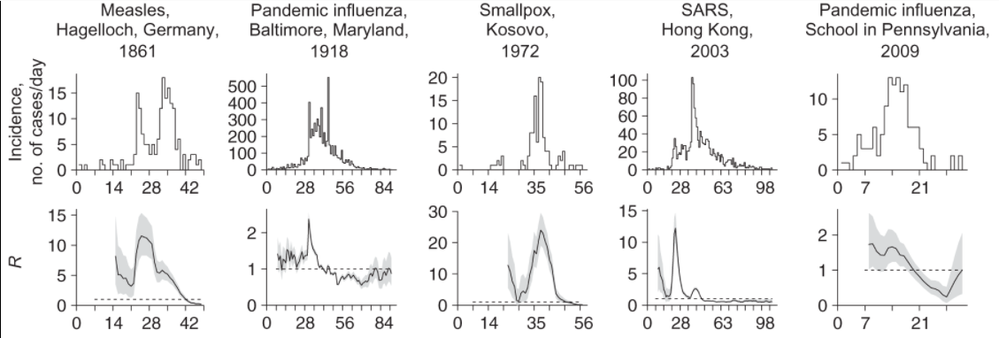
There's also a similar R package, EpiEstim, developed by a different group of researchers that has an Excel option. EpiEstim is based on a branching process model and estimates the R0 based on simple time series data. This model tries to capture the number of people each infected person will infect, but with an element of randomness (or stochasticity) -- like a random encounter in the grocery store with an infected person. The chart below (part of a larger graphic) shows the estimates of R0 generated with this model for five past outbreaks.
Sequencing Pathogen Genomes
"It shows novel characteristics. … It's Godzilla, King Kong and Frankenstein all in one."
- Contagion
Genetic analysis of SARS-CoV-2 samples from patients who become ill in different places and at different times can help researchers track the spread and mutation of the virus. This analysis can also help the quick identification of possible treatments. Researchers recently demonstrated a new machine learning approach to identifying the type of an unknown virus and its various strains based on its genome (i.e., figuring out its taxonomic classification, like the broad category of "coronavirus").
This approach converted the SARS-CoV-2 genomic sequence into a numerical representation (see the full research paper for details). The sequences of nearly 15,000 other viruses were also used in the training data. The researchers trained six different machine learning models (Linear Discriminant, Linear SVM, Quadratic SVM, Fine KNN, Subspace Discriminant, and Subspace KNN). The trained models then output their predictions for labels at the highest taxonomic level for the genomes of the virus strains causing COVID-19. The researchers then shifted the models to focus on the next, more specific taxonomic level and repeated the process again.
The graphic below shows the researchers' results from their last two tests, which classified 153 viral sequences into four sub-genera and COVID-19, and then classified 76 viral sequences either as other Sarbecovirus types or as COVID-19.
This strategy helped identify not only that SARS-CoV-2 should be correctly classified with other Coronaviridae and Betacoronavirus pathogens, but also that it has important similarities to other viruses found among bats. The researchers argue that their approach is faster (under 10 minutes, including 10-fold cross-validation) and can compare more, and more diverse, samples than previous analytic processes. While there may be insights here for the current pandemic, this kind of approach could be helpful in future outbreaks.
Predicting Disease Spread Despite Uncertainty
"And from there, using our model, based on the R0 of 3.2 … here is where we expect to be in 48 hours."
- Contagion
One epidemiological modeling approach is to create a "SIR model," which divides the population as a whole into "compartments": those who are susceptible to the disease, infected, and removed (either recovered from the illness and presumably granted some degree of immunity, or no longer in the population because they died).
However, generating such a model is never easy. Outbreak data can be impossible to gather accurately for many reasons, including institutional barriers, lack of testing, unknown or asymptomatic cases, and so forth. And, as we've seen, governments have implemented varied social distancing and isolation measures at different times, which can have unpredictable results on the number of people in that "susceptible" compartment.
To deal with all the sources of uncertainty, one group of researchers developed what they call an "eSIR" model -- an extended model including "a time-varying probability that a susceptible person meets an infected person or vice versa," as well as a new compartment to include people who are susceptible but choose self-quarantine. Both of these factors would vary in specific regions based on when isolation protocols were implemented.
To further incorporate uncertainty into the model, the researchers used a Markov chain Monte Carlo (MCMC) algorithm. (Here are two explanations for MCMC: one simpler, one more complex.) The MCMC approach allows for the approximation of a distribution that can't be directly known -- like the true number of SARS-CoV-2 infections -- or that is too expensive to figure out. The eSIR model's forecasts aim to reveal "turning points" in the outbreak. Turning points include when the daily number of new cases stops growing, and when the number of infected cases reaches its highest point. The model can also provide an estimate of the R0.
The researchers are distributing an R package called eSIR that generates the models, ggplot2 objects, and summary statistics. What's useful about this approach is that it can help determine which quarantine strategies might be most useful and when. As the researchers say, "... too strict quarantine can cause backfire; people may lose their trust and patience in their committed system, and consequently may try to reduce compliance or even avoid quarantine." The risk of implementing strict quarantine systems for prolonged periods has to be weighed against the gains in disease prevention. This model provides one approach to that important calculation.
Takeaways for All Modeling
What lessons can we learn from these still-preliminary models that extend beyond the pandemic? There are a few key items.
First, these models show rapid innovation to address an extremely complex situation. (The genomic and 'eSIR' studies described above are still "preprints," meaning they have not yet been peer-reviewed and have been released as quickly as possible to contribute to the scientific community's effort to address the pandemic.) It's truly impressive to see the creativity that researchers are applying to this crisis with such speed despite great stress, and it's inspiring for everyone as we seek solutions to many new challenges.
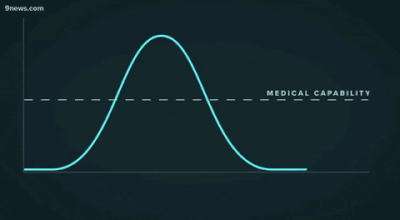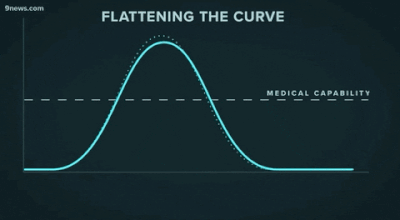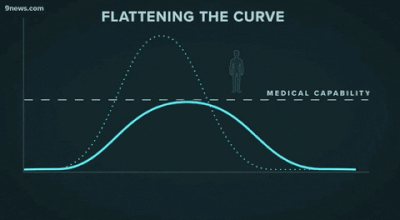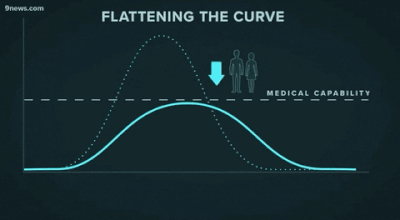
Second, another challenge that may be familiar to data people: getting decisionmakers to take action based on insights from data. The great "flattening the curve" visualization and related forecasting seem to have had a strong impact on policymakers and the general public. Likewise, data experts need to be able to communicate about their analyses and models clearly -- for example, with effective data visualizations -- and organizations should build data literacy across all areas.
Finally, the research I reviewed frequently mentioned the challenge of getting quality COVID-19 data upon which to build good models. Even in normal times, it can be hard to get the kind and quality of data we need. Models for aspects of the pandemic -- or any other phenomenon -- are only as good as the data upon which they are built. Data abounds everywhere, but not all are trustworthy, usable, or relevant. Every organization needs solid data gathering, management, and analytics structures in place (as these epidemiologists recommend for outbreaks). With that preparation, if a crisis of any kind emerges, your responses can be informed by accurate, relevant, up-to-date data readily at hand.
Original. Reposted with permission.
Bio: Susan Currie Sivek, Ph.D., is a writer and data geek who enjoys figuring out how to explain complicated ideas in everyday language, sometimes in silly ways. She appreciates good food, science fiction, and dogs.
Related: