Become a Pro at Pandas, Python’s Data Manipulation Library
Pandas is one of the most popular Python libraries for cleaning, transforming, manipulating and analyzing data. Learn how to efficiently handle large amounts of data using Pandas.
By Julien Kervizic, Senior Enterprise Data Architect at GrandVision NV
The Pandas library is the most popular data manipulation library for Python. It provides an easy way to manipulate data through its data-frame api, inspired from R’s data-frames.

Understanding the Pandas library
One of the keys to getting a good understanding of Pandas is to understand that Pandas is mostly a wrapper around a series of other Python libraries. The main ones being Numpy, SQLAlchemy, Matplotlib and openpyxl.
The core internal model of the data-frame is a series of numpy arrays, and Pandas functions such as the now deprecated “as_matrix” return results in that internal representation.
Pandas leverages other libraries to get data in and out of data-frames, SQLAlchemy, for instance, is used through the read_sql and to_sql functions. And openpyxl and xlsx writer are used for read_excel and to_excel functions.
Matplotlib and Seaborn are instead used to provide an easy interface to plot information available within a data frame, using command such as df.plot().
Numpy’s Pandas — Efficient Pandas
One of the complain that you often hear is that Python is slow or that it is difficult to handle large amount of data. Most often than not, this is due to poor efficiency of the code being written. It is true that native Python code tends to be slower than compiled code, but libraries like Pandas effectively provides an interface in Python code to compiled code. Knowing how to properly interface with it, let us get the best out of Pandas/Python.
APPLY VECTORIZED OPERATIONS
Pandas, like its underlying library Numpy, performs vectorized operations more efficiently than performing loops. These efficiencies are due to vectorized operations being performed through C compiled code, rather than native Python code and on the ability of vectorized operations to operate on entire datasets.
The apply interface allows to gain some of the efficiency by using a CPython interfaces to do the looping:
df.apply(lambda x: x['col_a'] * x['col_b'], axis=1)
But most of the performance gain would be obtained from the use of vectorized operations themselves, be it directly in Pandas or by calling its internal Numpy arrays directly.

As you can see from the picture above the difference in performance can be drastic, between processing it with a vectorized operation (3.53ms) and looping with Apply to do an addition (27.8s). Additional efficiencies can be obtained by directly invoking the numpy’s arrays and API, eg:

Swifter: Swifter is a Python library that makes it easy to vectorize different types of operations on dataframe, its API is fairly similar to that of the Apply function.
EFFICIENT DATA STORING THROUGH DTYPES
When loading a data-frame into memory, be it through read_csv, or read_excel or some other data-frame read function, SQL makes type inference which might prove to be inefficient. These APIs allow you to specify the types of each columns explicitly. This allows for a more efficient storage of data in memory.
df.astype({'testColumn': str, 'testCountCol': float})
Dtypes are native object from Numpy, which allows you to define the exact type and number of bits used to store certain informations.
Numpy’s dtype np.dtype('int32') would for instance represent a 32 bits long integer. Pandas default to 64 bits integer, we could save half the space by using 32 bits:
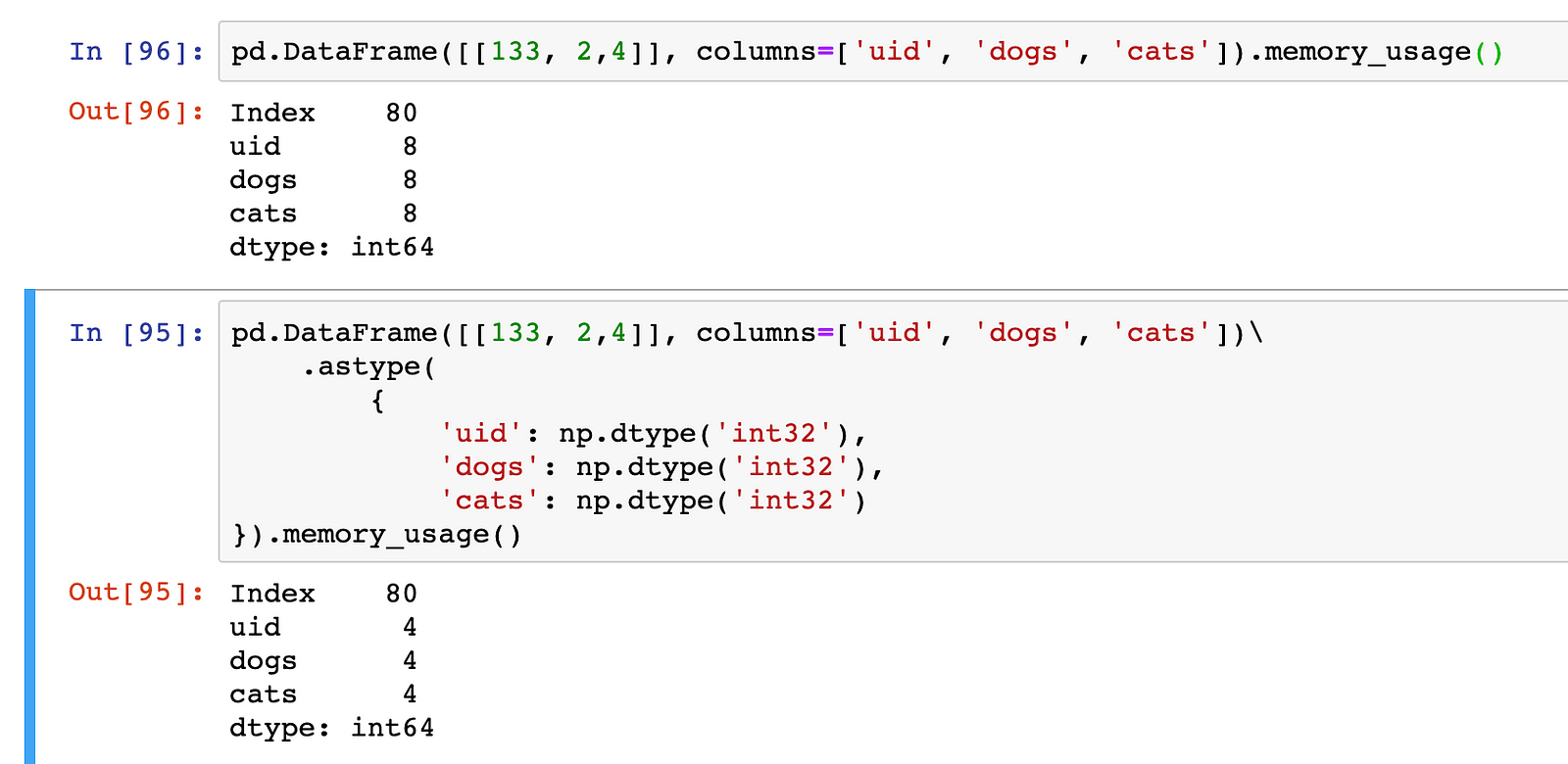
memory_usage() shows the number of bytes used by each of the columns, since there is only one entry (row) per column, the size of each int64 column is 8 bytes and of int32 4 bytes.
Pandas also introduces the categorical dtype, that allows for efficient memory utilization for frequently occurring values. In the example below, we can see a 28x decrease in memory utilization for the field posting_date when we converted it to a categorical value.
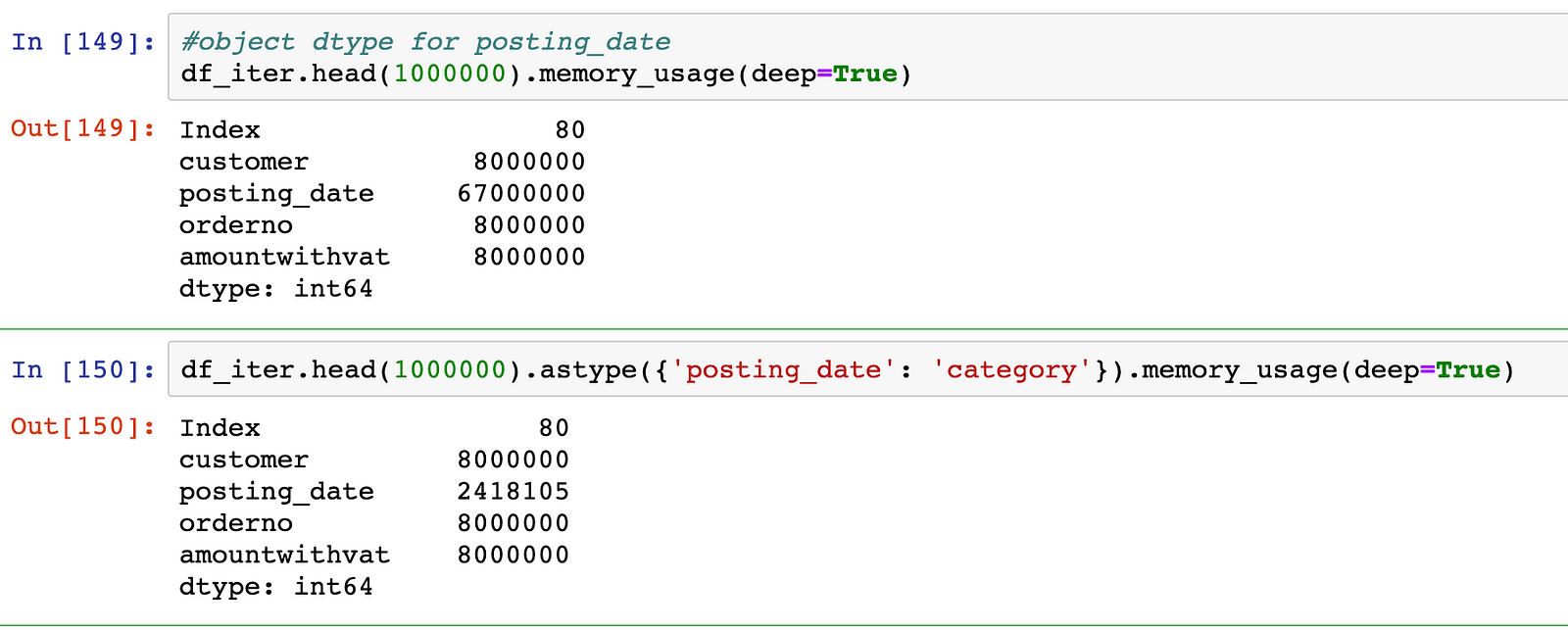
In our example, the overall size of the data-frame drops by more than 3X by just changing this data type:
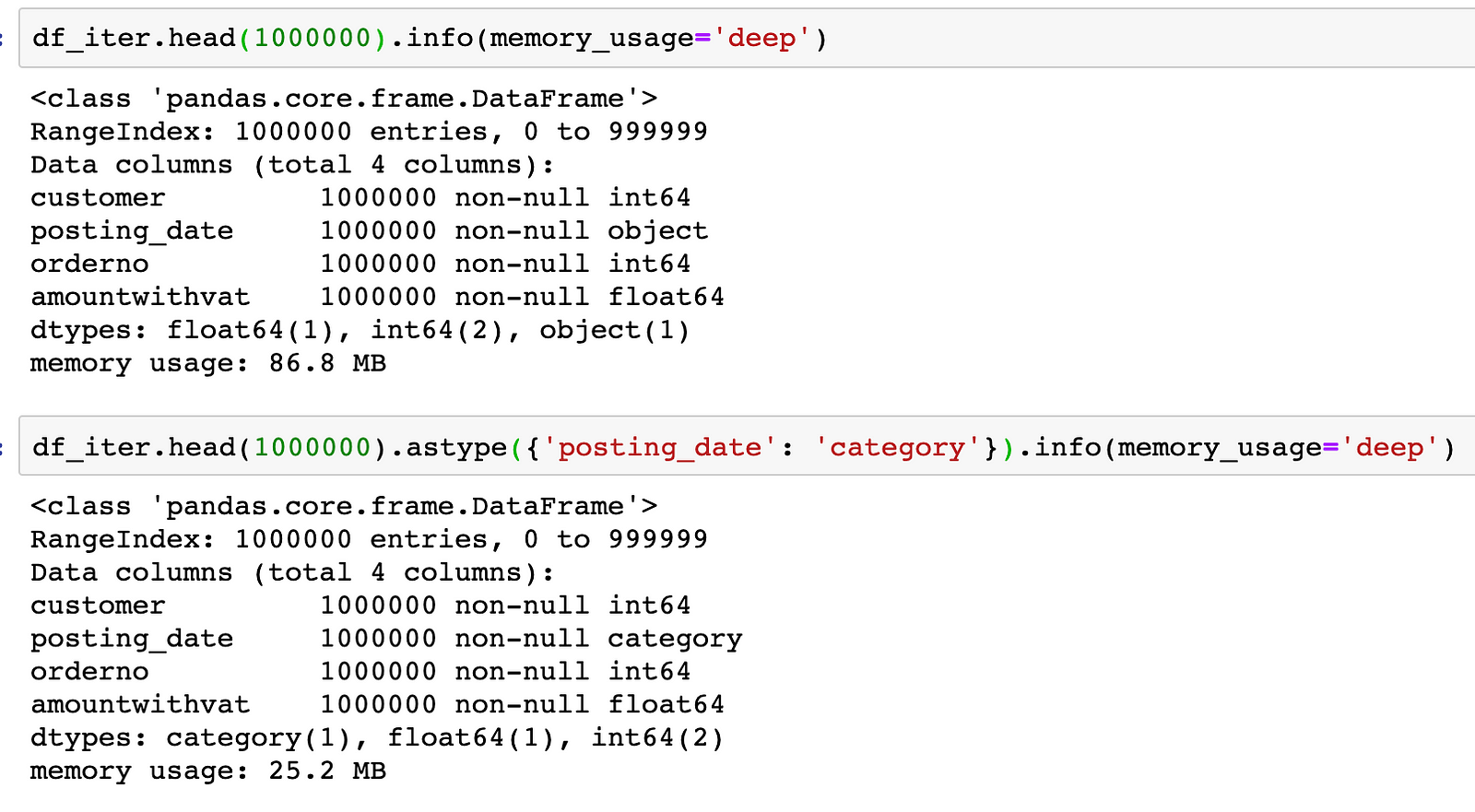
Not only using the right dtypes allows you to handle larger datasets in memory, it also makes some computations become more effective. In the example below, we can see that using categorical type brought a 3X speed improvement for the groupby/sum operation.
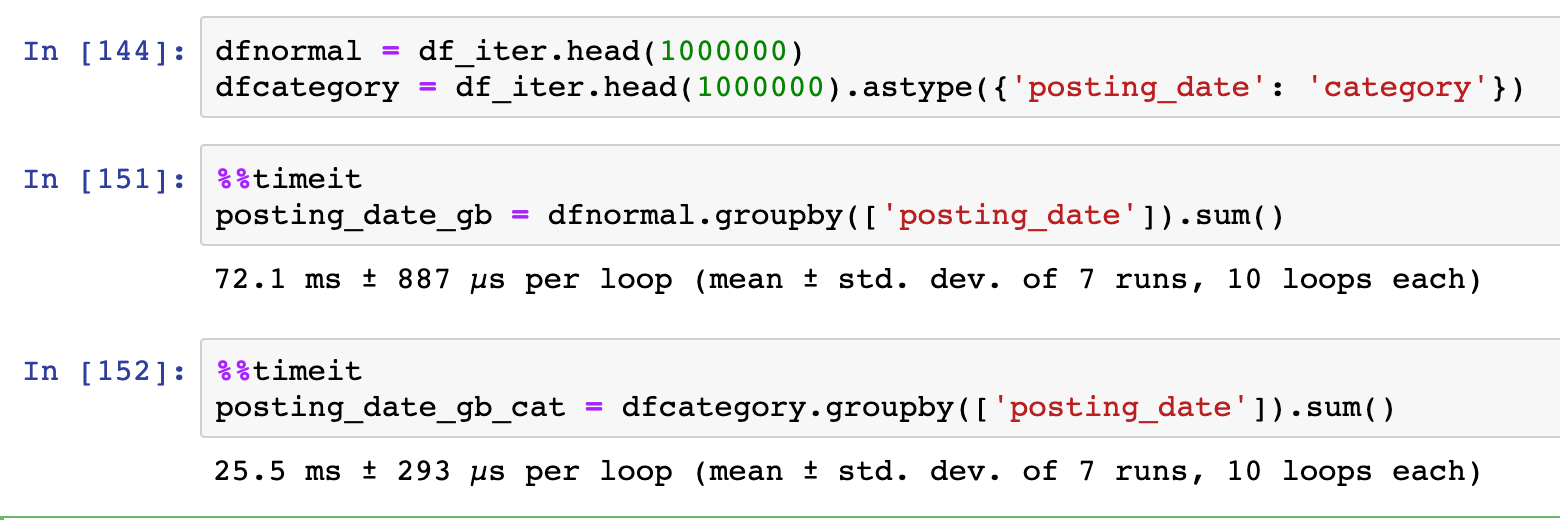
Within Pandas, you can define the dtypes during the data load (read_ ) or as a type conversion (astype).
CyberPandas: CyberPandas is one of the different library extensions that enables a richer variety of datatypes by supporting ipv4 and ipv6 data types and storing them efficiently.
HANDLING LARGE DATASETS WITH CHUNKS
Pandas allows for the loading of data in a data-frame by chunks, it is therefore possible to process data-frames as iterators and be able to handle data-frames larger than the available memory.
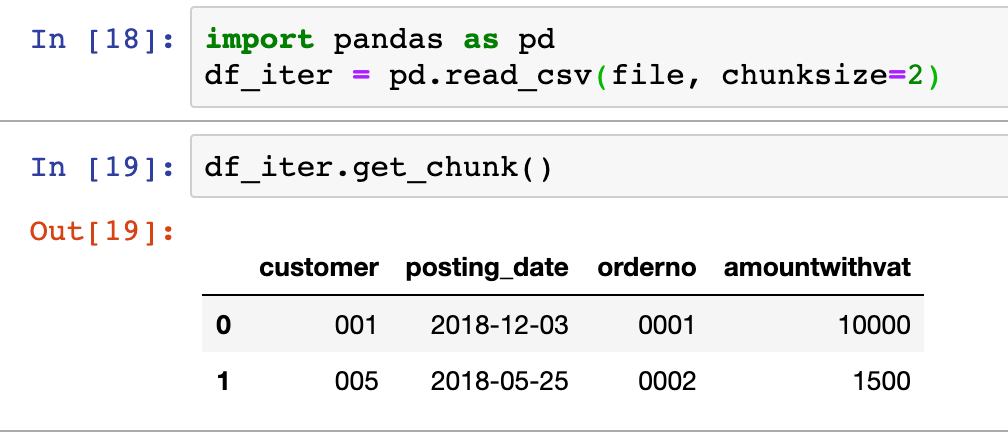
The combination of defining a chunksize when reading a data source and the get_chunk method, allows Pandas to process data as an iterator. For instance, in the example shown above, the data frame is read 2 rows at the time. These chunks can then be iterated through:
i = 0
for a in df_iter:
# do some processing chunk = df_iter.get_chunk()
i += 1
new_chunk = chunk.apply(lambda x: do_something(x), axis=1)
new_chunk.to_csv("chunk_output_%i.csv" % i )
The output of which can then be fed to a csv file, pickled, exported to a database, etc…
setting up a operator by chunks also allows certain operations to be performed through multi-processing.
Dask: It is a framework built on top of Pandas and built with multi-processing and distributed processing in mind. It makes use of collections of chunks of Pandas data-frames both in memory and on disk.
SQL Alchemy’s Pandas — Database Pandas
Pandas also is built up on top of SQLAlchemy to interface with databases, as such it is able to download datasets from diverse SQL type of databases as well as push records to it. Using the SQLAlchemy interface ( rather than using the Pandas API) directly allows us to do certain operations not natively supported within Pandas such as transactions or upserts:
SQL TRANSACTIONS
Pandas can also make use of SQL transactions, handling commits and rollbacks. Pedro Capelastegui, explained in one of his blog posts, how Pandas could take advantage of transactions through a SQLAlchemy context manager.
with engine.begin() as conn:
df.to_sql(
tableName,
con=conn,
...
)
The advantage of using a SQL transaction, is the fact that the transaction would roll back, if the data load fail.
SQL Extension
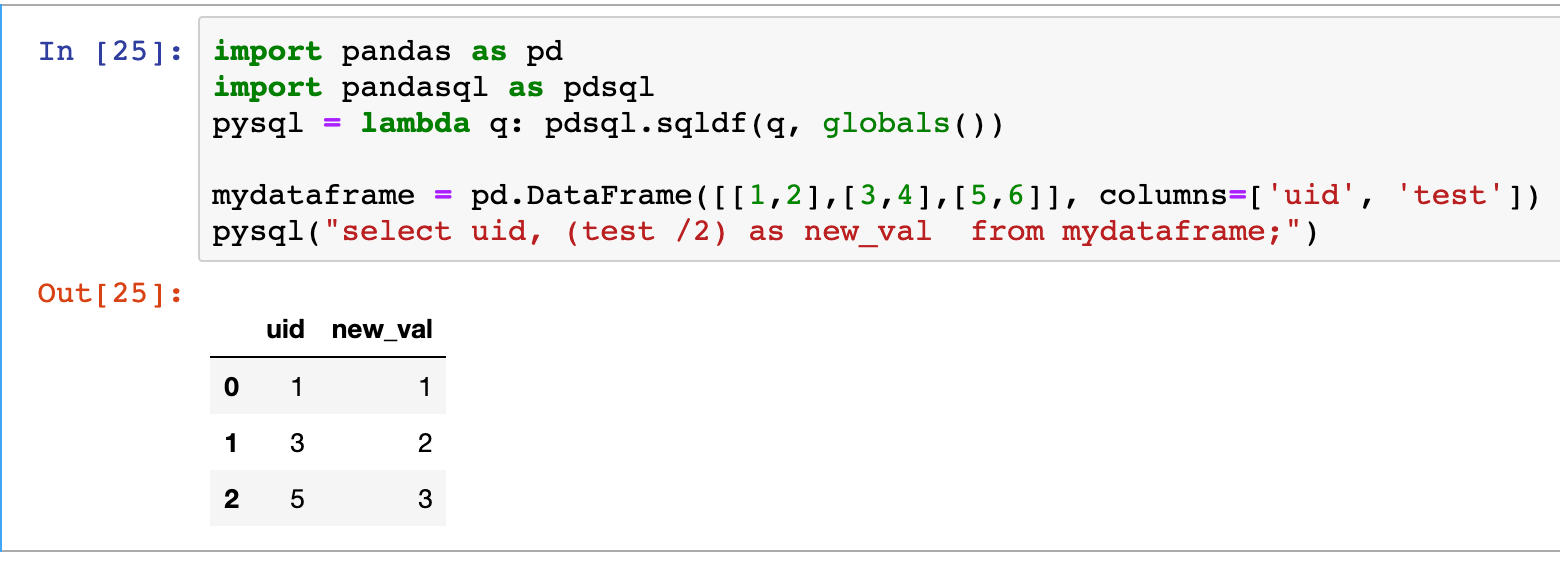
PandaSQL
Pandas has a few SQL extension such as pandasql a library that allows to perform SQL queries on top of data-frames. Through pandasql the data-frame object can be queried directly as if they were database tables.
SQL UPSERTs
Pandas doesn’t natively support upsert exports to SQL on databases supporting this function. Patches to Pandas exists to allow this feature.
MatplotLib/Seaborn — Visual Pandas
Matplotlib and Seaborn visualization are already integrated in some of the dataframe APIs such as through the .plot command. There is a fairly comprehensive documentation as how the interface works, on Pandas website.
Extensions: Different extensions exists such as Bokeh and plotly to provide interactive visualization within Jupyter notebooks, while it is also possible to extend matplotlib to handle 3D graphs.
Other Extensions
Quite a few other extensions for Pandas exists, which are there to handle no-core functionalities. One of them is tqdm, which provides a progress bar functionality for certain operations, another is pretty Pandas which allows to format dataframes and add summary informations.
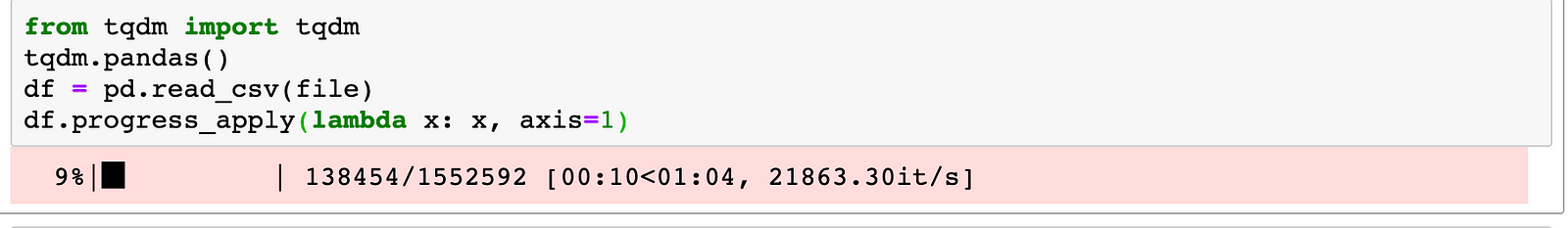
tqdm
tqdm is a progress bar extension in Python that interacts with Pandas, it allows user to see the progress of maps and apply operations on Pandas dataframe when using the relevant function (progress_map and progress_apply):
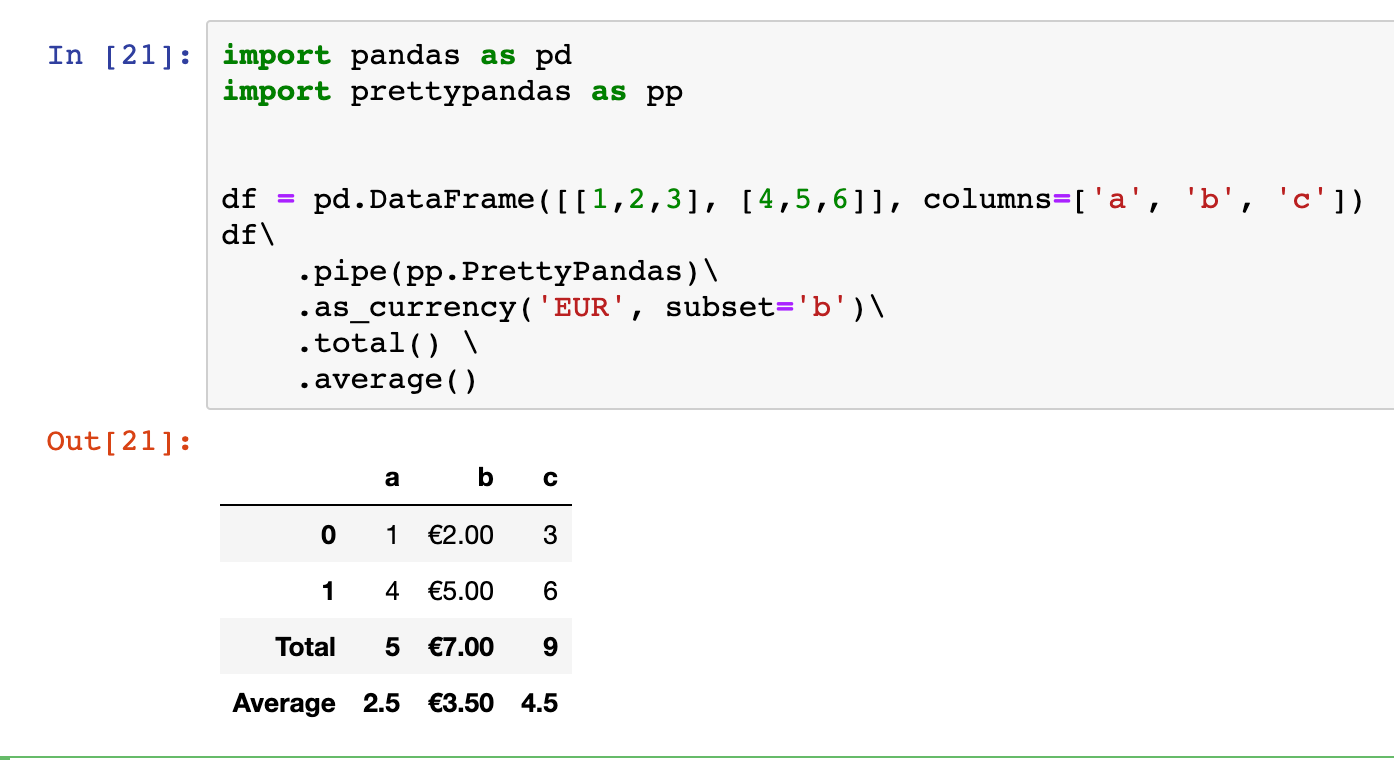
PrettyPandas
PrettyPandas is a library that provides an easy way to format data-frames and to add table summaries to them:
Bio: Julien Kervizic is a Senior Enterprise Data Architect at GrandVision NV.
Original. Reposted with permission.
Related:
