Iterative Initial Centroid Search via Sampling for k-Means Clustering
Thinking about ways to find a better set of initial centroid positions is a valid approach to optimizing the k-means clustering process. This post outlines just such an approach.
In this post, we will look at using an iterative approach to searching for a better set of initial centroids for k-means clustering, and will do so by performing this process on a sample of our full dataset.
What do we mean by "better?" Since k-means clustering aims to converge on an optimal set of cluster centers (centroids) and cluster membership based on distance from these centroids via successive iterations, it is intuitive that the more optimal the positioning of these initial centroids, the fewer iterations of the k-means clustering algorithms will be required for convergence. Therefore, thinking about ways to find a better set of initial centroid positions is a valid approach to optimizing the k-means clustering process.
What we will do differently, specifically, is to draw a sample of data from our full dataset, and run short runs of of the k-means clustering algorithm on it (not to convergence), short runs which will include, out of necessity, the centroid initialization process. We will repeat these short runs with a number of randomly initialized centroids, and will track the improvement to the measurement metric -- within-cluster sum-of-squares -- for determining goodness of cluster membership (or, at least, one of the valid metrics for measuring this). The final centroids associated with the random centroid initialization iteration process which provide the lowest inertia is the set of centroids which we will carry forward to our full dataset clustering process.
The hope is that this up-front work will lead to a better set of initial centroids for our full clustering process, and, hence, a lesser number of k-means clustering iterations and, ultimately, less time required to fully cluster a dataset.
This would obviously not be the only method of optimizing centroid initialization. In the past we have discussed the naive sharding centroid initialization method, a deterministic method for optimal centroid initialization. Other forms of modifications to the k-means clustering algorithm take different approaches to this problem as well (see k-means++ for comparison).
This post will approach our task as follows:
- prepare the data
- prepare our sample
- perform centroid initialization search iterations to determine our "best" collection of initial centroids
- use results to perform clustering on full dataset
A future post will perform and report comparisons of results between various approaches to centroid initialization, for a more comprehensive understanding of the practicalities of implementation. For now, however, let's introduce and explore this particular approach to centroid initialization.
Preparing the data
For this overview, we will use the 3D road network dataset.
Since this particular dataset has no missing values, and no class labels, our data preparation will primarily constitute normalization, along with dropping a column which identifies a geographical location from where the additional 3 columns worth of measurements come from, which is not useful for our task. See the dataset description for additional details.
import numpy as np
import pandas as pd
from sklearn import preprocessing
# Read dataset
data = pd.read_csv('3D_spatial_network.csv', header=None)
# Drop first column (not required)
data.drop(labels=0, axis=1, inplace=True)
# Normalize data (min/max scaling)
data_arr = data.values
sc = preprocessing.MinMaxScaler()
data_sc = sc.fit_transform(data_arr)
data = pd.DataFrame(data_sc)
Let's check out a sampling of our data:
data.sample(10)
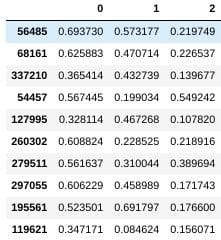
Preparing the sample
Next, we will pull our sample that will be used to find our "best" initial centroids. Let's be clear about exactly what we are doing:
- we are pulling a single set of samples from our dataset
- we will then perform successive rounds k-means clustering on this sample data, each iteration of which will:
- randomly initialize k centroids and perform n iterations of the k-means clustering algorithm
- the initial inertia (within-cluster sum-of-squares) of each centroid will be noted, as will its final inertia, and the initial centroids which provide the greatest increase in inertia over n iterations will be chosen as our initial centroids for the full dataset clustering
- we will then perform full k-means clustering on the full dataset, using the initial clusters found in the previous step
2 important points:
- Why not use greatest decrease in inertia? (The hope being that the initial momentum in this area will continue.) Doing so would also be a valid choice to explore (and changing a single line of code would allow for this). An arbitrary initial experimentation choice, and one which could use more investigation. However, the repetitive execution and comparison on a number of samples initially showed that the lowest inertia and greatest decrease in inertia coincide a great majority of the time, and so the decision may, in fact, be arbitrary, but also inconsequential in practice.
- Of particular clarification, we are not sampling multiple times from our dataset (e.g. once from our dataset for each iteration of centroid initialization). We are sampling once for all iterations of a single centroid initialization search. One sample, from which we will randomly derive initial centroids many times. Contrast this with the idea of repetitive sampling, once for each centroid initialization iteration.
Below, we set:
- sample size as a ratio of our full dataset
- random state for reproducibility
- number of clusters (k) for our dataset
- number of iterations (n) for our k-means algorithm
- number of attempts at finding our best chance initial centroids while clustering on our sample dataset
We then set our sample data
# Some variables SAMPLE_SIZE = 0.1 RANDOM_STATE = 42 NUM_CLUSTERS = 10 # k NUM_ITER = 3 # n NUM_ATTEMPTS = 5 # m data_sample = data.sample(frac=SAMPLE_SIZE, random_state=RANDOM_STATE, replace=False) data_sample.shape
Now that we have our data sample (data_sample) we are ready to perform iterations of centroid initialization for comparison and selection.
Clustering the sample data
Since Scikit-learn's k-means clustering implementation does not allow for easily obtaining centroids between clustering iterations, we have to hack the workflow a bit. While the verbose option does output some useful information in this regard directly to screen, and redirecting this output and then post-parsing it would be one approach to getting what we need, what we will do is write our own outer iteration loop to control for our n variable ourselves.
This means that we need to count iterations and capture what we need between these iterations, after each clustering step has run. We will then wrap that clustering iteration loop in a centroid initialization loop, which will initialize k centroids from our sample data m times. This is the hyperparameter specific to our particular instantiation of the k-means centroid initialization process, beyond "regular" k-means.
Given our above parameters, we will be clustering our dataset into 10 clusters (NUM_CLUSTERS, or k), we will run our centroid search for 3 iterations (NUM_ITER, or n), and we will attempt this with 5 random initial centroids (NUM_ATTEMPTS, or m), after which we will determine our "best" set of centroids to initialize with for full clustering (in our case, the metric is the lowest within-cluster sum-of-squares, or inertia).
Prior to any clustering, however, let's see what a sample of what a single initialization of our k-means looks like, prior to any clustering iterations.
from sklearn.cluster import KMeans
km = KMeans(n_clusters=NUM_CLUSTERS, init='random', max_iter=1, n_init=1)#, verbose=1)
km.fit(data_sample)
print('Pre-clustering metrics')
print('----------------------')
print('Inertia:', km.inertia_)
print('Centroids:', km.cluster_centers_)
Pre-clustering metrics ---------------------- Inertia: 898.5527121490726 Centroids: [[0.42360342 0.20208702 0.26294088] [0.56835267 0.34756347 0.14179924] [0.66005691 0.73147524 0.38203476] [0.23935675 0.08942105 0.11727529] [0.58630271 0.23417288 0.45793108] [0.1982982 0.11219503 0.23924021] [0.79313864 0.52773534 0.1334036 ] [0.54442269 0.60599501 0.17600424] [0.14588389 0.29821987 0.18053109] [0.73877864 0.8379479 0.12567452]]
In the code below, note that we have to manually track our centroids at the start of each iteration and at the end of each iteration, given that we are managing these successive iterations ourselves. We then feed these end centroids into our next loop iteration as the initial centroids, and run for one iteration. A bit tedious, and aggravating that we can't get this out of Scikit-learn's implementation directly, but not difficult.
final_cents = []
final_inert = []
for sample in range(NUM_ATTEMPTS):
print('\nCentroid attempt: ', sample)
km = KMeans(n_clusters=NUM_CLUSTERS, init='random', max_iter=1, n_init=1)#, verbose=1)
km.fit(data_sample)
inertia_start = km.inertia_
intertia_end = 0
cents = km.cluster_centers_
for iter in range(NUM_ITER):
km = KMeans(n_clusters=NUM_CLUSTERS, init=cents, max_iter=1, n_init=1)
km.fit(data_sample)
print('Iteration: ', iter)
print('Inertia:', km.inertia_)
print('Centroids:', km.cluster_centers_)
inertia_end = km.inertia_
cents = km.cluster_centers_
final_cents.append(cents)
final_inert.append(inertia_end)
print('Difference between initial and final inertia: ', inertia_start-inertia_end)
Centroid attempt: 0 Iteration: 0 Inertia: 885.1279991728289 Centroids: [[0.67629991 0.54950506 0.14924545] [0.78911957 0.97469266 0.09090362] [0.61465665 0.32348368 0.11496346] [0.73784495 0.83111278 0.11263995] [0.34518925 0.37622882 0.1508636 ] [0.18220657 0.18489484 0.19303869] [0.55688642 0.35810877 0.32704852] [0.6884195 0.65798194 0.48258798] [0.62945726 0.73950354 0.21866185] [0.52282355 0.12252092 0.36251485]] Iteration: 1 Inertia: 861.7158412685387 Centroids: [[0.67039882 0.55769658 0.15204125] [0.78156936 0.96504069 0.09821352] [0.61009844 0.33444322 0.11527662] [0.75151713 0.79798919 0.1225065 ] [0.33091899 0.39011157 0.14788905] [0.18246521 0.18602087 0.19239602] [0.55246091 0.3507018 0.33212609] [0.68998302 0.65595219 0.48521344] [0.60291234 0.73999001 0.23322449] [0.51953015 0.12140833 0.34820443]] Iteration: 2 Inertia: 839.2470653106332 Centroids: [[0.65447477 0.55594052 0.15747416] [0.77412386 0.952986 0.10887517] [0.60761544 0.34326727 0.11544127] [0.77183027 0.76936972 0.12249837] [0.32151587 0.39281244 0.14797103] [0.18240552 0.18375276 0.19278224] [0.55052636 0.34639191 0.33667632] [0.691699 0.65507199 0.48648245] [0.59408317 0.73763362 0.23387334] [0.51879974 0.11982321 0.34035345]] Difference between initial and final inertia: 99.6102464383905 ...
After this is done, let's see how we did in our centroid search. First we check a list of our final inertias (or within-cluster sum-of-squares), looking for the lowest value. We then set the associated centroids as the initial centroids for our next step.
# Get best centroids to use for full clustering best_cents = final_cents[final_inert.index(min(final_inert))] best_cents
And here's what those centroids look like:
array([[0.55053207, 0.16588572, 0.44981164],
[0.78661867, 0.77450779, 0.11764745],
[0.656176 , 0.55398196, 0.4748823 ],
[0.17621429, 0.13463117, 0.17132811],
[0.63702675, 0.14021011, 0.18632431],
[0.60838757, 0.39809226, 0.14491584],
[0.43593405, 0.49377153, 0.14018223],
[0.16800744, 0.34174697, 0.19503396],
[0.40169376, 0.15386471, 0.23633233],
[0.62151433, 0.72434071, 0.25946183]])
Running full k-means clustering
And now, with our best initial centroids in hand, we can run k-means clustering on our full dataset. As Scikit-learn allows us to pass in a set of initial centroids, we can exploit this by the comparatively straightforward lines below.
km_full = KMeans(n_clusters=NUM_CLUSTERS, init=best_cents, max_iter=100, verbose=1, n_init=1) km_full.fit(data)
This particular run of k-means converged in 13 iterations:
... start iteration done sorting end inner loop Iteration 13, inertia 7492.170210199639 center shift 1.475641e-03 within tolerance 4.019354e-06
For comparison, here's a full k-means clustering run using only randomly-initialized centroids ("regular" k-means):
km_naive = KMeans(n_clusters=NUM_CLUSTERS, init='random', max_iter=100, verbose=1, n_init=1) km_naive.fit(data)
This run took 39 iterations, with a nearly-identical inertia:
... start iteration done sorting end inner loop Iteration 39, inertia 7473.495361902045 center shift 1.948248e-03 within tolerance 4.019354e-06
I leave finding the difference of execution times between the 13 iterations and 39 iterations (or similar) to the reader. Needless to say, eating up a few cycles ahead of time on a sample of data (in our case, 10% of the full dataset) saved considerable cycles in the long run, without sacrifice to our overall clustering metric.
Of course, additional testing before drawing any generalizations is warranted, and in a future post I will run some experiments on a number of centroid initializiation methods on a variety of datasets and compare with some additional metrics, hopefully to get a clearer picture of ways to go about optimizing unsupervised learning workflows.
Related:
- Toward Increased k-means Clustering Efficiency with the Naive Sharding Centroid Initialization Method
- Comparing Distance Measurements with Python and SciPy
- Machine Learning Workflows in Python from Scratch Part 2: k-means Clustering
